
Detection of CDR3s diversity and its prediction of persistent high-risk HPV infection and cervical intraepithelial neoplasia risk: a prospective study
Introduction
The incidence and prevalence of cervical cancer have been decreasing year by year with the maturity of three-stage cervical cancer screening and the concept of screening and treatment (1) and follow-up (2). Advances in cervical cancer screening and vaccination have exacerbated geographic differences, including in the United States and China (3,4). In 2016, the American College of Obstetricians and Gynecologists still recommended immediate surgical treatment for cervical intraepithelial neoplasia (CIN) grade 2 (CIN 2) at ≥25 years old, and all CIN 3 or more severe (≥ CIN 3) (5). However, previous studies have confirmed that most CIN 2 will naturally return to normal, not all CIN 3 will progress to cervical cancer, and the probability that untreated CIN 3 progressing to cervical cancer in 30 years is still less than 50% (6,7). In addition, HPV mRNA and DNA (Hybrid Capture 2, HC2) assays are highly sensitive for CIN 2 or more severe (≥ CIN 2) and much more sensitive than cytology, while HPV mRNA is more specific than HC2 and similar to cytology (8). Therefore, current screening guidelines inevitably lead to overtreatment. More importantly, excessive examination and treatment bring extravagant psychological burden (9) and economic pressure to patients.
Sustained infection of HPV will lead to precancerous lesions, and the precancerous lesions progressing to invasive cancer is relevant to cell immunity of B cells and T cells (10-14). To a large extent, complementarity-determining region 3s (CDR3s) diversity of T cell receptor (TCR) repertoire is able to represent the capacity of T cell immune diversity (15,16). Previous studies have found that TCR CDR3s immune diversity may be an effective indicator for predicting persistent high-risk HPV (hr-HPV) infection and CIN risk (17,18).
We performed this prospective study to simulate the natural pathogenesis of cervical cancer, and used the quantitative index of immune diversity, namely diversity 50 (D50), to determine the potential role of CDR3s. According to follow up, the relationship between CDR3s immune diversity and natural clearance or persistent infection of HPV was reported. At the same time, the information of low-grade CIN degradation, persistence and progression was collected to seek the relationship between CDR3s immune diversity and CIN 1 risk. We present the following article in accordance with the TREND reporting checklist (available at https://gpm.amegroups.com/article/view/10.21037/gpm-21-48/rc).
Methods
Study population and procedure
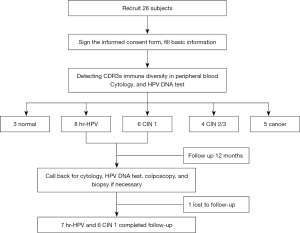
This single center study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by Ethics Committee of West China Second University Hospital, Sichuan University, China (No. 2013-036) and informed consent was taken from all individual participants. The subjects were recruited from October 2016 to December 2017. The exclusion criteria: (I) women during pregnancy; (II) history of cervical surgery; (III) congenital or primary immunodeficiency disease; (IV) long-term medication history of glucocorticoids, immunosuppressants, immunomodulators and so on; (V) history of mental illness; (VI) history of other malignant diseases, radiotherapy and chemotherapy; (VII) anemia, malnutrition; (VIII) history of bone marrow, thymus, thyroid and other diseases; (IX) history of diabetes, or kidney transplantation; (X) smokers, or drinkers. The inclusion criteria were: (I) females aged 30–64 years; (II) being sexually active but non-pregnant; (III) no history of cervical surgery; (IV) having civil decision-making authority and signed informed consent. Finally, 3 normal controls, 8 with hr-HPV infections, 6 with CIN 1, 4 with CIN 2/3, and 5 with cervical cancer were included. In this prospective single-center study, peripheral blood, cervical exfoliated cells and frozen cervical tissues samples were collected. Figure 1 shows the procedure flowchart for our study.
T cell separation
Ten ml of peripheral blood was put into the tube with heparin and T cell isolation was performed by magnetic activated cell sorting (Miltenyi Biotec, Germany). CD8+ T cells were first screened negatively by anti-CD4 magnetic beads, and then separated by anti-CD8 magnetic beads for positive screening. CD4+ T cells were isolated by positive screening using anti-CD4 magnetic beads. CD4+ CD25+ Tr cells were positively screened by anti-CD25 magnetic beads. All extracted cells were immediately stored in RNA preservation solution (Qiagen, Germany).
Amplicon rescued multiplex polymerase chain reaction (ARM-PCR)
RNA was extracted by using the RNeasy Mini Kit (Qiagen, Germany), and the complementary DNA (Toyobo, Japan) was subsequently used to perform real-time PCR with SYBR™ chemistry (Takara, China) using gene-specific primers (forward 5'-GGCAGAGGAGAGGAGTACCA-3'; reverse 5'-AGCCATAGCCACCTACTCCA-3'). The amplified products from different samples were mixed and loaded into 2% agarose. By electrophoresis, DNA fragment from agarose gel with purified for a 250–500 bp DNA fragments.
Second generation high throughput sequencing (HTS)
Titanium reagent and 454 GS FLX system were used to sequence DNA (SeqWright, USA). The sequencing results of each test sample result were classified according to the coding tags. The reference sequences of cell lines V and J were downloaded from the IMGT database and read by Irmap. CDR3s border area was read through the surveying and mapping information mapping sequence. CDR3s confined region was isolated and transformed into amino acid sequence.
The diversity index is defined as 100 area under the curve between percentage of total reads and percentage of unique CDR3s, when the frequencies of unique CDRs are accumulated from most frequent to least frequent. Importantly, the D50 value is used to evaluate the diversity of immune database, and it explains at least 50% of CDR3s with a minimum proportion of differential CDR3s (19,20). D50 detection abandons the complex model fitting and only focuses on richer subgroups, calibrated by the area of the whole population (21). D50 value of a specific repertoire is positively related to CDR3s sequences diversity, whereas negatively the cumulative frequency difference between each CDR3s clone (21,22).
HPV DNA test
All the cervical exfoliated cells samples were detected by LUMIX system (Tellgen, China), which can qualitatively distinguish the 9 genotypes of hr-HPV (31, 35, 39, 45, 51, 56, 59, 66 and 68), and specificity detect hr-HPV subtype of 16, 18, 33, 52 and 58 closely associated with cervical cancer.
ThinPrep cytology test (TCT)
The cervical exfoliated cells samples were tested by the method ThinPrep (Hologiccytyc, USA). Cytological diagnosis was precisely made by two cytologists accordance with the term of the Bethesda system.
Colposcopy, cervical biopsy and pathological diagnosis
Colposcopy was performed by a professional gynecologist and the cervical biopsy was taken if necessary. All specimens were routinely embedded in paraffin and sectioned in the pathology department of West China Second Hospital. First, the two histopathologists read the slides independently. When the results were inconsistent, the third pathologist read the slides again. The final diagnosis was made by majority rule. Surgical treatment was indicated for women with histological diagnosis of ≥ CIN 2.
Follow-up
The subjects of the hr-HPV group and the CIN 1 group were respectively followed up to confirm the HPV infection, cytology and the status of cervix in the 12th month.
Statistical analysis
These analyses and charting were performed using GraphPad Prism software (version 7.00) and SPSS (version 24.0). All experiments were strictly carried out with the instructions and repeated at least three times. For quantitative data, t-test/variance analysis/nonparametric test were used for comparison; for qualitative data, chi square test was used. P<0.05 was statistically significant.
Results
The description of patients’ characteristics at the baseline year
The grouping depended on the results of colposcopy-directed biopsy, which was only but one diagnosed chronic inflammation in the hr-HPV group. The characteristics of these groups were presented in Table 1. There was no significant difference of the age, menarche, menstrual conditions, contraceptives, pregnant frequency, parturition frequency and D50 value among the five groups. All the subjects had no smoking and drinking history. More importantly, there was significant among age at sex debut (P=0.002). In the cancer group, the main clinic manifestations were neoplasm and bleeding with all HPV positive results.
Table 1
| Variables | Total (n=26) | Normal (n=3) | hr-HPV (n=8) | CIN 1 (n=6) | CIN 2/3 (n=4) | Cancer (n=5) | P |
|---|---|---|---|---|---|---|---|
| Age (year) | 45.7±9.7 | 38±5.3 | 51.0±8.6 | 47.8±7.5 | 36.5±7.9 | 46.8±11.6 | 0.069 |
| Menarche (year) | 14.6±2.0 | 15.3±0.6 | 15.4±2.4 | 14.5±1.2 | 14.3±3.3 | 13.3±1.0 | 0.510 |
| Age at sex debut (year) | 21.9±2.6 | 24.7±1.5 | 20.8±1.7 | 20.2±1.2 | 23.0±3.0 | 26.0±2.8 | 0.002 |
| Menstrual conditions | 0.129 | ||||||
| Premenopausal | 18 (69.2) | 3 (100.0) | 5 (62.5) | 2 (33.3) | 4 (100.0) | 4 (80.0) | |
| Postmenopausal | 8 (30.8) | 0 (0.0) | 3 (37.5) | 4 (66.7) | 0 (0.0) | 1 (20.0) | |
| Contraceptives | 0.173 | ||||||
| No | 11 (44.0) | 2 (66.7) | 2 (25.0) | 5 (83.3) | 1 (25.0) | 1 (25.0) | |
| Yes | 14 (56.0) | 1 (33.3) | 6 (75.0) | 1 (16.7) | 3 (75.0) | 3 (75.0) | |
| Pregnant frequency (times) | 0.279 | ||||||
| 0–1 | 6 (23.1) | 1 (33.3) | 1 (12.5) | 0 (0.0) | 2 (50.0) | 2 (40.0) | |
| 2–3 | 10 (38.5) | 1 (33.3) | 2 (25.0) | 5 (83.3) | 1 (25.0) | 1 (20.0) | |
| 4–5 | 10 (38.5) | 1 (33.3) | 5 (62.5) | 1 (16.7) | 1 (25.0) | 2 (40.0) | |
| Parturition frequency | 0.345 | ||||||
| 0–1 | 18 (69.2) | 3 (100.0) | 4 (50.0) | 4 (66.7) | 4 (100.0) | 3 (60.0) | |
| ≥2 | 8 (30.8) | 0 (0.0) | 4 (50.0) | 2 (33.3) | 0 (0.0) | 2 (40.0) |
Data are shown as mean ± standard deviation or number (percentage). hr-HPV, high-risk human papillomavirus; CIN, cervical intraepithelial neoplasia.
At baseline year, CDR3s diversity was not associated with cervical lesions
The prevalence of CDR3s diversity staining was 9.2±7.9, 5.7±5.6, 4.0±6.0, 13.6±7.7, 8.0±7.6 among women with normal, hr-HPV positive, CIN 1, CIN 2/3 and cancer biopsies. However, it was no statistically significant difference (P=0.252) (Table 2 and Table S1).
Table 2
| Group | D50 |
|---|---|
| Control (n=3) | 9.2±7.9 |
| hr-HPV (n=8) | 5.7±5.6 |
| CIN 1 (n=6) | 4.0±6.0 |
| CIN 2/3 (n=4) | 13.6±7.7 |
| Cancer (n=5) | 8.0±7.6 |
| P | 0.252 |
Data are shown as mean ± standard deviation. CDR3s, complementarity-determining region 3s; D50, diversity 50; hr-HPV, high-risk human papillomavirus; CIN, cervical intraepithelial neoplasia.
Based on the risk of cervical diseases, subjects were divided into the CIN less severe than CIN 2 (< CIN 2) group (normal, hr-HPV and CIN 1, n=17) and the ≥ CIN 2 group (CIN 2/3 and cancer, n=9). The CDR3s diversity between the two groups was not significant (P=0.093) (Table 3).
Table 3
| Cervical diseases | D50 |
|---|---|
| < CIN 2 (n=14) | 5.7±6.0 |
| ≥ CIN 2 (n=9) | 10.5±7.7 |
| P | 0.093 |
Data are shown as mean ± standard deviation. CDR3s, complementarity-determining region 3s; D50, diversity 50; CIN, cervical intraepithelial neoplasia.
At baseline screening, CDR3s diversity was not also associated with hr-HPV infection
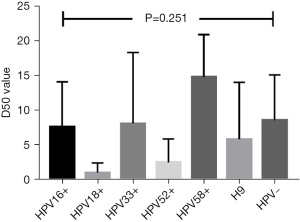
The D50 value was not associated with the HPV infected results (P=0.709) (Table 4). Further analysis of HPV subtypes also showed no statistical differences (P=0.251) (Figure 2).
Table 4
| HPV infection | D50 |
|---|---|
| HPV negative (n=4) | 8.6±6.6 |
| HPV positive (n=22) | 7.1±7.1 |
| P | 0.709 |
Data are shown as mean ± standard deviation. CDR3s, complementarity-determining region 3s; HPV, human papillomavirus; D50, diversity 50.
The CDR3s was unsignificant with the CIN1 regression and stabilization after 12 months
One patient in the hr-HPV group was lost to follow up. All results of visual inspection of the cervix were normal, including one Nessler’s cyst and one mild erosion. Based on the examination results of the baseline year and following up 12 months, the subjects were divided into the stabilization group and the degradation group.
According to the 12th-month HPV results, there was not apparent significance of the D50 value in the degradation group and the stabilization group (P=0.173) (Table 5). Only 1 case of atypical squamous cells of undetermined significance (ASC-US) was found and her biopsy result was chronic cervical and endocervical inflammation with squamous metaplasia. The rest results were normal in the CIN 1 group. It made statistical analysis impossible between the degradation group and the stabilization group in the CIN 1 group because of the too few cases (Table 6). Colposcopy examination was normal in patients with abnormal results from TCT or HPV test, according to cervical cancer screening guidelines.
Table 5
| hr-HPV | D50 |
|---|---|
| Stabilization (n=4) | 2.5±3.4 |
| Degradation (n=3) | 8.8±7.1 |
| P | 0.173 |
Data are shown as mean ± standard deviation. CDR3s, complementarity-determining region 3s; hr-HPV, high-risk human papillomavirus.
Table 6
| CIN 1 | D50 |
|---|---|
| Stabilization (n=1) | 0.84 |
| Degradation (n=5) | 4.7±6.5 |
| P | – |
Data are shown as mean ± standard deviation. CDR3s, complementarity-determining region 3s; TCT, ThinPrep cytology test; CIN, cervical intraepithelial neoplasia; D50, diversity 50.
Discussion
In this study, we used D50 to detect CDR3s for the first time, and explored the clinical value of CDR3s may work as a biomarker to predict the risk of persistent hr-HPV infection and CIN 1. We hypothesized that the higher the D50 value of CDR3s immune diversity detections, the greater the possibility of HPV natural clearance and CIN conversion to normal. In order to increase the external validity of the results, 5 groups of research subjects were selected to simulate the natural pathogenesis of cervical cancer. To minimize the bias of different diagnostic classifications and reduce misdiagnosis, pathologists independently finished the diagnosis of cytology and histopathology.
The results of the baseline study suggested that the differences of CDR3s between the < CIN 2 group and the ≥ CIN 2 group were not statistically significant. Previous studies have found that patients with CD4+ and CD8+ T cells in CIN 2/3 have a higher CIN clearance rate, especially eliminating HPV infected cervical epithelial cells (23-27). The diversity of TCR is mainly determined by CDR3s (15,16,21). But our results don’t support the unique study reporting the diversity of TCR CDR3s decreased during the cervix carcinogenesis and progression (17).
Therefore, the population in this prospective study included hr-HPV infected and CIN 1 women. The diagnosis of CIN 1 was determined after a thorough discussion by two or three histopathologists. The follow-up personnel did not know the CDR3s test results of HPV-positive and CIN 1 populations at baseline screening. In order to avoid destroying the natural course of cervical diseases, our team followed up on the target population using TCT and HPV test after 12 months.
For the CIN 1 group, all follow-ups were completed. Follow-up results revealed that 5 had normal cytology, and 1 had ASC-US without abnormalities of a referral to colposcopy. All the subjects did not find CIN 1 or more severe lesions. However, high TCR diversity may not degrade CIN 1 to normal, because only one patient with low D50 level persisted CIN 1, failing to statistical analysis. While for the CIN 1 persisted or progressed, Woo et al. reported that the infiltrating CD8+ T cells decrease activity (28). These findings suggest that CIN 1 women with low CDR3s diversity should receive close follow-up every 6 months or annually.
In the hr-HPV group, seven patients were followed up, of which 4 continued to be infected with HPV, and 3 became negative. The statistical results of only a few enrolled subjects suggested that there was no statistical difference of CDR3s between the two groups, though D50 in the hr-HPV degradation group was higher than that in the hr-HPV stabilization group. CDR3s cannot yet be considered as a biomarker to predict HPV-positive outcomes.
Our study has several limitations. First of all, this disease usually takes years for HPV infection to induce abnormal cell transformation and then for the lesion to appear (29). Considering with our 12-month follow-up, expanding the sample size of enrolled subjects and extending the follow-up time are inevitable choices to more effectively determine the predictive value of CDR3s in future studies. Secondly, some of the subjects’ CIN 1 lesions may be small at baseline screening, or all of the CIN 1 lesions’ tissues may have been removed at biopsy, which may also interfere with the follow-up results. In addition, the confounding factors may affect the outcome of D50 and the cervical lesions, including HLA alleles and haplotypes (30). For example, there was only one statistically significant result of age at sex debut but it is unclear whether this is clinically significant there were low n values and no apparent correlation with level of disease. It’s necessary to amplify samples number.
Due to the limitation of research funding, it is too relatively small of the sample size to find out the difference of CDR3s. Thus, CDR3s is not yet considered as a biomarker to predict HPV-positive outcomes. The detection of CDR3s may assist in the clinical management of CIN 1. Women with CIN 1 and decrease of CDR3s diversity may benefit from closer follow-up at frequently intervals.
Acknowledgments
Funding: This work was supported by grants from the National Natural Science Foundation of China (No. 81602504).
Footnote
Reporting Checklist: The authors have completed the TREND reporting checklist. Available at https://gpm.amegroups.com/article/view/10.21037/gpm-21-48/rc
Data Sharing Statement: Available at https://gpm.amegroups.com/article/view/10.21037/gpm-21-48/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://gpm.amegroups.com/article/view/10.21037/gpm-21-48/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by Ethics Committee of West China Second University Hospital (No. 2013-036) and informed consent was taken from all individual participants.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- World Health Organization. WHO Guidelines for Screening and Treatment of Precancerous Lesions for Cervical Cancer Prevention. Geneva: World Health Organization Press; 2013.
- Massad LS, Einstein MH, Huh WK, et al. 2012 updated consensus guidelines for the management of abnormal cervical cancer screening tests and cancer precursors. Obstet Gynecol 2013;121:829-46. [Crossref] [PubMed]
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin 2020;70:7-30. [Crossref] [PubMed]
- Liao G, Jiang X, She B, et al. Multi-Infection Patterns and Co-infection Preference of 27 Human Papillomavirus Types Among 137,943 Gynecological Outpatients Across China. Front Oncol 2020;10:449. [Crossref] [PubMed]
- Practice Bulletin No. 157. American College of Obstetricians and Gynecologists. Obstet Gynecol 2016;127:e1-20.
- Schiffman M, Rodríguez AC. Heterogeneity in CIN3 diagnosis. Lancet Oncol 2008;9:404-6. [Crossref] [PubMed]
- McCredie MR, Sharples KJ, Paul C, et al. Natural history of cervical neoplasia and risk of invasive cancer in women with cervical intraepithelial neoplasia 3: a retrospective cohort study. Lancet Oncol 2008;9:425-34. [Crossref] [PubMed]
- Monsonego J, Hudgens MG, Zerat L, et al. Evaluation of oncogenic human papillomavirus RNA and DNA tests with liquid-based cytology in primary cervical cancer screening: the FASE study. Int J Cancer 2011;129:691-701. [Crossref] [PubMed]
- Khan MJ, Werner CL, Darragh TM, et al. ASCCP Colposcopy Standards: Role of Colposcopy, Benefits, Potential Harms, and Terminology for Colposcopic Practice. J Low Genit Tract Dis 2017;21:223-9. [Crossref] [PubMed]
- den Boon JA, Pyeon D, Wang SS, et al. Molecular transitions from papillomavirus infection to cervical precancer and cancer: Role of stromal estrogen receptor signaling. Proc Natl Acad Sci U S A 2015;112:E3255-64. [Crossref] [PubMed]
- Kurmyshkina OV, Kovchur PI, Schegoleva LV, et al. T- and NK-cell populations with regulatory phenotype and markers of apoptosis in circulating lymphocytes of patients with CIN3 or microcarcinoma of the cervix: evidence for potential mechanisms of immune suppression. Infect Agent Cancer 2017;12:56. [Crossref] [PubMed]
- Guzmán-Olea E, Bermúdez-Morales VH, Peralta-Zaragoza O, et al. Molecular Mechanism and Potential Targets for Blocking HPV-Induced Lesion Development. J Oncol 2012;2012:278312. [Crossref] [PubMed]
- Sheu BC, Chang WC, Lin HH, et al. Immune concept of human papillomaviruses and related antigens in local cancer milieu of human cervical neoplasia. J Obstet Gynaecol Res 2007;33:103-13. [Crossref] [PubMed]
- Nguyen HH, Broker TR, Chow LT, et al. Immune responses to human papillomavirus in genital tract of women with cervical cancer. Gynecol Oncol 2005;96:452-61. [Crossref] [PubMed]
- Marrack P, Kappler J. Positive selection of thymocytes bearing alpha beta T cell receptors. Curr Opin Immunol 1997;9:250-5. [Crossref] [PubMed]
- Miqueu P, Guillet M, Degauque N, et al. Statistical analysis of CDR3 length distributions for the assessment of T and B cell repertoire biases. Mol Immunol 2007;44:1057-64. [Crossref] [PubMed]
- Cui JH, Lin KR, Yuan SH, et al. TCR Repertoire as a Novel Indicator for Immune Monitoring and Prognosis Assessment of Patients With Cervical Cancer. Front Immunol 2018;9:2729. [Crossref] [PubMed]
- Lang Kuhs KA, Lin SW, Hua X, et al. T cell receptor repertoire among women who cleared and failed to clear cervical human papillomavirus infection: An exploratory proof-of-principle study. PLoS One 2018;13:e0178167. [Crossref] [PubMed]
- Kuo HC, Pan CT, Huang YH, et al. Global Investigation of Immune Repertoire Suggests Kawasaki Disease Has Infectious Cause. Circ J 2019;83:2070-8. [Crossref] [PubMed]
- Wu J, Liu D, Tu W, et al. T-cell receptor diversity is selectively skewed in T-cell populations of patients with Wiskott-Aldrich syndrome. J Allergy Clin Immunol 2015;135:209-16. [Crossref] [PubMed]
- Hou D, Ying T, Wang L, et al. Immune Repertoire Diversity Correlated with Mortality in Avian Influenza A (H7N9) Virus Infected Patients. Sci Rep 2016;6:33843. [Crossref] [PubMed]
- Guo C, Wang Q, Cao X, et al. High-Throughput Sequencing Reveals Immunological Characteristics of the TRB-/IgH-CDR3 Region of Umbilical Cord Blood. J Pediatr 2016;176:69-78.e1. [Crossref] [PubMed]
- Origoni M, Parma M, Dell'Antonio G, et al. Prognostic significance of immunohistochemical phenotypes in patients treated for high-grade cervical intraepithelial neoplasia. Biomed Res Int 2013;2013:831907. [Crossref] [PubMed]
- Monnier-Benoit S, Mauny F, Riethmuller D, et al. Immunohistochemical analysis of CD4+ and CD8+ T-cell subsets in high risk human papillomavirus-associated pre-malignant and malignant lesions of the uterine cervix. Gynecol Oncol 2006;102:22-31. [Crossref] [PubMed]
- Maskey N, Thapa N, Maharjan M, et al. Infiltrating CD4 and CD8 lymphocytes in HPV infected uterine cervical milieu. Cancer Manag Res 2019;11:7647-55. [Crossref] [PubMed]
- Piersma SJ, Jordanova ES, van Poelgeest MI, et al. High number of intraepithelial CD8+ tumor-infiltrating lymphocytes is associated with the absence of lymph node metastases in patients with large early-stage cervical cancer. Cancer Res 2007;67:354-61. [Crossref] [PubMed]
- Trimble CL, Clark RA, Thoburn C, et al. Human papillomavirus 16-associated cervical intraepithelial neoplasia in humans excludes CD8 T cells from dysplastic epithelium. J Immunol 2010;185:7107-14. [Crossref] [PubMed]
- Woo YL, Sterling J, Damay I, et al. Characterising the local immune responses in cervical intraepithelial neoplasia: a cross-sectional and longitudinal analysis. BJOG 2008;115:1616-21; discussion 1621-2. [Crossref] [PubMed]
- Molling JW, de Gruijl TD, Glim J, et al. CD4(+)CD25hi regulatory T-cell frequency correlates with persistence of human papillomavirus type 16 and T helper cell responses in patients with cervical intraepithelial neoplasia. Int J Cancer 2007;121:1749-55. [Crossref] [PubMed]
- Alifu M, Fan P, Kuerban G, et al. Frequency distribution of HLA alleles and haplotypes in Uyghur women with advanced squamous cell cervical cancer and relation to HPV status and clinical outcome. Arch Gynecol Obstet 2018;297:757-66. [Crossref] [PubMed]
Cite this article as: Tang D, Liao GD, Cao HY, Kang LN, Wei BB, Xi MR, Zeng X, Chen MY. Detection of CDR3s diversity and its prediction of persistent high-risk HPV infection and cervical intraepithelial neoplasia risk: a prospective study. Gynecol Pelvic Med 2022;5:22.


